Moreover, Genes whose relative transcription ranges showed a fold transform FC 1. 33 and SAM |Score | two had been consid ered for being up regulated, and those with an FC 0. 75 and SAM |Score | two have been viewed as to be down regu lated. In this research, 575 transcripts showed a amount of expression that differed considerably from that in the manage group with H. parasuis serovar five SH0165 strain contaminated group. These incorporated 428 genes annotated with DAVID or by looking against the GenBank database. Amid these, 338 genes have been up regulated and 90 genes had been down regulated. The functions from the DE genes had been analyzed by Mole cular Annotation Procedure 3. 0 application. While in the MAS 3. 0 device, the GO terms and KEGG pathways are ranked with statistical significance by calculating their p values based on hypergeometric distribution.
Go terms and KEGG pathways with selelck kinase inhibitor p values less than 0. 05 are considered statistically signifi cant. Among the 428 annotated genes, a complete of 229 genes have been grouped into 156 classes depending on biological course of action Gene Ontology terms with all the p values less than 0. 05. Several GO terms were associated using the immune process. These had been inflammatory response. immune response. complement activation, classical pathway and leukocyte migration. Especially, the DE genes linked with immune and inflammatory response suggested that they perform roles in host defense response to H. parasuis infection. To gain insight into the diverse biological processes of H. parasuis infec tion, a pathway examination by KEGG database was per formed on DE genes.
The substantial pathways selleck chemicals MG-132 largely contained cell adhesion molecules, cytokine cytokine receptor interaction, complement and coagulation cascades, toll like receptor signaling pathway, MAPK signaling pathway, which advised that the host took diverse approaches to activate immune and inflammatory response upon H. parasuis infection. Validation of microarray information by quantitative real time PCR So that you can verify the statistical significance of our findings, we performed quantitative serious time PCR analysis of the related genes in our original samples utilized in microarray study. Eleven genes had been chosen for qPCR evaluation. 10 picked genes that were up regulated in microarray also showed drastically greater expression in H. parasuis serovar five infected sam ples than during the handle samples established by qPCR analysis.
The ppp1r13l gene that was down regulated in microarray information also showed drastically decrease expres sion in H. parasuis serovar five infected samples than inside the manage samples by qPCR. STRING evaluation on the relationships concerning DE genes STRING 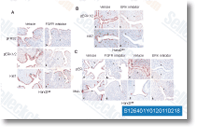 is a web based mostly interface that could predict pro tein associations which may imply direct physical binding and may also indicate indirect interaction such as participa tion during the very same metabolic pathway or cellular procedure within the basis of genomic context, large throughput experiments, co expression and data in the literature genes had been analyzed employing STRING for predicting network of proteins encoded by DE genes.
is a web based mostly interface that could predict pro tein associations which may imply direct physical binding and may also indicate indirect interaction such as participa tion during the very same metabolic pathway or cellular procedure within the basis of genomic context, large throughput experiments, co expression and data in the literature genes had been analyzed employing STRING for predicting network of proteins encoded by DE genes.
Monthly Archives: July 2014
Biological replicates were pooled for making repre sentative samp
Biological replicates were pooled to produce repre sentative samples for deep sequencing evaluation. Throughout the two groups of triplicate information, after normalization from the generated 95 bp PE raw reads, 15,683,828, 13,040,780 and 25,660,654 reads have been obtained from C1 C3, and sixteen,306,312, 15,589,848 and ten,906,906 reads from V1 V3, respectively. To assess the high-quality of se quencing, the reads had been mapped for the zebrafish refer ence genome. Through the reads of each group, productive mapping occurred for ten,823,266, 9,584,828, 18,321,987, 12,209,418, 11,675,593 and eight,605,104 reads. However, 4,860,562, three,455,952 and 7,338,667 unmapped reads had been produced from C1 C3, although four,096,894, 3,914,255 and two,301,802 unmapped reads had been uncovered in V1 V3, respectively.
we approach to con duct de novo analysis of those unmapped reads to gener ate a better reference of immune relevant genes in zebrafish. Examination of differential expression among WED and mock immunized zebrafish liver To determine the differentially expressed genes, the tran scriptome information of zebrafish liver from two days following WED immunization and mock immunization have been ana INK1197 ic50 lyzed by using the DESeq package deal in R application. The criteria of a two fold or greater adjust in expres sion and p value 0. 05 were selected to determine drastically up regulated or down regulated genes following immunization. The magnitude distribution of your appreciably transformed genes was illu strated by MA plot analysis. Making use of these cri teria, a complete of 4565 genes have been considerably differentially expressed higher than two fold, like 2186 up regulated genes and 2379 down regulated genes.
Annotation of Anacetrapib the differentially expressed genes was attained as a result of BLASTN similar ity searches towards the Ensembl zebrafish RefSeq mRNA database. To carry out an unbiased annotation on the functions in the differentially expressed genes identified by DESeq examination, GO examination of differentially expressed genes was carried out by two bioinformatics tools, DAVID and BiNGO plugin. Between the 4565 dif ferentially expressed genes, DAVID presented functional annotation for 3891 genes. GO annotated differentially expressed genes mainly belonged on the three practical clusters, and had been distributed among greater than 70 categories. The differentially expressed genes during the cluster of biological processes had been uncovered to become largely related to stimulus response, immune response, regulation of immune process process, and regulation of growth procedure. To recognize the biological pathways which can be active within the zebrafish in the early stages of WED immunization, 4565 differentially expressed genes had been mapped to ca nonical signaling pathways uncovered in KEGG.
These lupus nephritis genes were further shown to not be associ
These lupus nephritis genes were further shown to not be associated with the normal ageing process based on the observed differences between healthy young and old C57BL6 mice. A broad range of biological functions was represented among the lupus nephritis genes identified in this study. As expected, given the loss of kidney function, the vast majority of genes involved in metabolic pathways are down regulated in nephritis and, given the inflammatory nature of the disease, many of the signalling pathway genes are up regulated. Glomerular disease is a sig nificant component in lupus nephritis. A recent study identified a glomerulus enriched gene set. We used data from this study to determine if the nephritis associated genes are enriched in the glomerular gene set.
We found a highly signif icant over representation of the glomerular genes consistent with glomerular involvement. A recent study by Liu and colleagues reported on 126 nephri tis associated genes in the MRLlpr model. Of these, 37 genes were present in the nephritis signature reported here. Commonalities were noted in the nephritis signatures between these two models, selleck inhibitor such as the antigen presentation and complement pathways as well as various IFN regulated genes and immunoglobulins. A good overlap was also noted between our mouse nephritis gene set and 68 human lupus nephritis genes derived from laser captured glomeruli from SLE patients. Additional similarities may be present, but probably lie outside the statistical parameters of both datasets.
A profound normalisation of expression levels of lupus nephri tis genes was observed in mice treated with sirolimus, both for metabolic as well as signalling pathways. Affected metabolic pathways in lupus nephritis include fatty more bonuses acid degradation, gly colysis pathways and leucinevalineisoleucine degradation. Transcripts for BCKDHA and DBT, two enzymes in the branched chain amino acid metabolism pathway required for the catabolism of leucine, valine and isoleucine, are reduced in nephritis, perhaps leading to the accumulation of leucine in diseased tissue. Interestingly, leucine activates the target of sirolimus inhibition, mTOR, leading to increased protein syn thesis, and in addition we noted an increase in ribosomal RNA transcripts in the disease state. This physiological con nection suggests that mTOR pathway activation may be increased by leucine in disease, providing perhaps an addi tional mechanism for sirolimus efficacy. Levels of these tran scripts were returned to asymptomatic levels in sirolimus treated mice. Several genes in the mitochondrial electron transport chain are also down regulated in the disease state, and mitochondrial dysfunction has been implicated in kidney function impairment.
However, offered the lack of head to head data for direct compa
On the other hand, given the lack of head to head information for direct comparison, network meta analyses are crucial in order to calculate the expected efficacy of biologic DMARDs. Indirect comparisons of interventions might be manufactured as a result of a common comparator. Our objective was to execute a network meta evaluation for abatacept following a systematic review from the pub lished clinical proof of abatacept and all other exist ing biologic DMARDs on the market, licensed in Europe for patients that failed to react to MTX or within the system of getting this kind of a license. The aim of your review was to estimate the relative efficacy of abatacept in mixture with MTX in Overall health Evaluation Questionnaire change from baseline in contrast to other rele vant biologic DMARDs plus MTX while in the treatment method of sufferers with RA with inadequate response to MTX.
Like a secondary aim, we studied the efficacy in terms of response charges from the American College Rheumatology Criterion for 50% improvement and in Dis ease Exercise Score in 28 joints defined remis sion. Resources inhibitor MP-470 and methods Systematic evaluate A protocol was designed to define the search system and a systematic evaluate carried out consecutively to determine individuals randomised controlled trials, which investigated the efficacy of biologic DMARDs licensed to treat RA with insufficient response to at least 1 conventional DMARD. MEDLINE and EMBASE databases had been searched concurrently using Datastar. Even further searches have been undertaken to the Cochrane Library, the American College of Rheumatol ogy and European League Towards Rheumatism conferences, as well as the technology appraisals for your Uk.
Searches included a mixture of zero cost text and Medline Topic Headings terms for dis ease terms with drug names, and were limited to human RCTs published, in English, amongst January 1980 and January 2010. The systematic analysis ON-01910 Estybon was carried out by two research ers, with discussions among the two to come to agree ment in case of discrepancies. The total text content articles had been assessed for inclusion according to the following selec tion criteria treatment combinations of MTX with abatacept, adalimumab, certolizumab, etanercept, goli mumab, infliximab, rituximab or tocilizumab in compari son with each other or Placebo MTX. RA patients with an inadequate response or intolerance to former remedy with not less than 1 typical DMARD.
clinical endpoints of HAQ CFB,  American School of Rheumatology Criterion of 50% improvement and remission defined by a Sickness Activity Score including a 28 joint count significantly less than 2. six. at 24 andor 52 weeks. Information collection For each picked research, the details of style and design, choice criteria, research population characteristics, interventions, outcome measures, length of observe up and benefits were extracted and recorded in information extraction varieties.
American School of Rheumatology Criterion of 50% improvement and remission defined by a Sickness Activity Score including a 28 joint count significantly less than 2. six. at 24 andor 52 weeks. Information collection For each picked research, the details of style and design, choice criteria, research population characteristics, interventions, outcome measures, length of observe up and benefits were extracted and recorded in information extraction varieties.
Protein was quantified with BCA Protein assay kit, with BSA appli
Protein was quantified with BCA Protein assay kit, with BSA made use of being a traditional. Surface plasmon resonance measurements The binding kinetics and affinity of sdAbA1 for purified CypA protein were obtained using the ProteON XPR36 protein interaction array process. A GLC chip was activated, on which recombinant CypA protein was immobilized at 25 C during the vertical orientation. Following that, the remaining carb oxyl groups were blocked with one M ethanolamineHCl. Finally, sdAbA1 was injected during the hori zontal orientation at 5 concentrations in twofold serial dilution down from 24 nM to 1. 5 nM at a flow price of 50 ulminute. Antibody binding was evaluated by simultan eously flowing 6 sdAb concentrations that ranging from 0 to 24 nM more than the CypA coated chip for 180 seconds, and then monitoring dissociation for 720 seconds.
The data have been analyzed with ProteON Manager 3. 1 computer software, and binding constants selleck chemical had been established using a eleven Langmuir binding model. Peptidyl prolyl cistrans isomerase activity assay The peptidyl prolyl cistrans isomerase activity was established inside a coupled assay with chymotrypsin making use of the tetrapeptide substrate Suc Ala Phe Professional Phe p nitroanilide. All reagents had been pre equilibrated until finally the temperature reached 0 C. Within a one ml cuvette, purified CypA was mixed with 10 ul chymotrypsin, along with the volume was adjusted to 975 ul with assay buffer. The reaction was initiated by the addition of 25 ul tetrapeptide substrate at a last concentration of 37. five uM. Improvements in absorbance on account of released p nitroaniline had been measured each 10 seconds for any optimum of 250 seconds within a spectrophotometer at 390 nm.
Cleavage in the tetrapeptide CPI-613 substrate from the absence of CypA was applied as a blank con trol, along with the addition of CypA was used being a beneficial control for PPIase action. CypA was additional at a concentration of 5. 0 uM. The result of sdAbA1 for the PPIase action of CypA was performed together with the pre incubation of CypA with sdAbA1 for two hrs in advance of addition to the reaction system. Induction, remedy, and evaluation of collagen induced arthritis The collagen induced arthritis model was con structed as described previously. Male DBA1 mice have been immunized intradermally with the base within the tail with 200 ug chicken kind II collagen emulsified in Freunds full adjuvant on the age of 8 to 12 weeks.
On day 21 right after principal immunization, mice have been provided a booster injection intradermally working with the same concentra tion of sort II collagen but with Freunds incomplete adju vant. Given that disease onset within this model varies broadly for each mouse, and that not all mice build arthritis, we in complete used a hundred mice to construct the CIA model. When immunized mice commenced to demonstrate clinical signs and symptoms, 30 arthritic mice have been chosen to obtain distinctive remedies and have been randomly assigned in cages.
As shown in Figure 3D, salivary killing action was similar concer
As shown in Figure 3D, salivary killing activity was equivalent involving RA subjects and wholesome controls. Therefore, even though there have been greater colonization prices of Candida and decreased IL 17A dependent salivary AMPs, salivary candi dacidal function appeared to be preserved in RA topics, that’s constant with their clinical resistance to OPC. Discussion In this review, we found that PBMCs from RA sufferers showed impaired Candida induced IL 17A production, in spite of total elevated basal IL 17A production as well as a preserved capability of CD4 cells to differentiate in res ponse to Th17 differentiating cytokines in vitro. The impaired Candida exact response was associated with an greater rate of RA topics colonized with Candida also as diminished expression of BD2, an IL 17A depen dent salivary AMP.
Nevertheless, salivary killing action against Candida was preserved in RA topics. Therefore, while there may be clearly a trend in the direction of increased suscep tibility to C. albicans colonization in RA, much of your effector antifungal immune response is reversible p53 inhibitor retained, consist ent together with the clinical resistance to oropharyngeal candidia sis in RA individuals. Genome broad association examine data level to a function to the Th17IL 17 axis in RA, as risk alleles influence Th17 generation and maintenance, trafficking or IL 17A signal transduction. Clinically, active RA has become associ ated with elevated fractions of Th17 cells compared with wholesome controls, and men and women that respond to TNF inhibitors are reported to present reductions in Th17 cells in contrast with nonresponders.
Erosive arth ritis in many animal versions is IL 17A dependent, as treatment with blocking antibodies ameliorates dis ease, and disease induction is mild or absent selleck chemicals in IL 17A deficient mice. Therefore, agents that inhibit the Th17 pathway at many factors, such as inhibitors of JAK kinases, IL 23, IL 17A and IL 17RA, are now being used or evaluated in RA together with other autoimmune condi tions. Since nearly all the RA sufferers in this review had DMARD controlled illness, we employed this group as being a reference population. An acknowledged limitation is ex clusion of therapy na ve patients with poorly managed disease. A great adhere to up shall be to assess longitudinal pathogen particular responses, beginning prior to drug treat ment is initiated. Nevertheless, these findings are internally consistent and recapitulate the characteristic clinical phenotype of RA, wherever overt susceptibility to OPC is seldom observed.
Our findings also suggest there may well be a threshold impact of IL 17A in mediating host defense to Candida, exactly where even minimal quantities of IL 17A are adequate for protective immunity. Our locating that Th17 cells in RA subjects have been decreased relative to controls contrasts with some prior scientific studies,  but may be explained from the undeniable fact that these patients had controlled disease with an regular Condition Action Score of 3.
but may be explained from the undeniable fact that these patients had controlled disease with an regular Condition Action Score of 3.
These enzymes kind the initial line of defense against superoxide
These enzymes form the very first line of defense against superoxide and hydrogen peroxide. The resultant secondary oxidation solutions may nevertheless harm DNA, proteins and lipid, and require additional detoxification. This second line of defense against ROSlipid peroxidation is supplied by enzymes such as the GSTs. Various examples support a modifying impact of oxidative strain genes on the relationships involving dietary factors and breast cancer. We are going to summarize next the potential modifying roles of oxidative pressure genes on the relationships amongst n 3 fatty acids, ITCs, and tea and breast cancer. Glutathione S transferases GSTM1, GSTT1, and GSTP1 Glutathione related metabolism can be a important mechanism for cellular protection against agents that generate oxidative pressure, guarding cells against cytotoxic items of lipid peroxidation.
GSTs are induced selleck under situations of oxidative anxiety, and alpha. pi. mu. and theta class GSTs are active in detoxification of quite a few products, which includes reactive oxidant harm to DNA and lipids, for instance organic epoxides, lipid hydroperoxides, and unsaturated aldehydes. Folks lacking these enzymes might have decreased removal of lipid peroxidation items and, therefore, could practical experience higher cancer protection, as supported by results from our study. We located that women with genetic polymorphisms causing lower or no activity in detoxifying genes had much more protection from marine n 3 fatty acids than those with typical alleles, putatively mainly because more cytotoxic peroxidation agents could attain the cancerous and pre cancerous cells and lead to damage.
Recent final results on cyclin D1, an intracellular cell cycle regulatory protein with checkpoint function, and breast cancer risk further assistance our hypo thesis of an oxidative anxiety induced apoptosis mechanism underlying the diet plan induced breast cancer protection. CCND1 has been shown to modulate selleckchem growth arrest and apoptosis following exposure to ionizing radiation oxidative DNA harm. Within a recent study, the protective impact of your heterozygous CCND1 GA genotype on breast cancer danger was restricted to conditions of elevated oxidative anxiety, characterized by absence of antioxidant GSTM1 and GSTT1 enzymes. Also, CCND1 also interacted with marine n 3 and n six fatty acids to influence breast cancer risk, using the protection being restricted to females having a higher intake degree of n six fatty acids, or possibly a low intake amount of marine n three fatty acids. Similarly, ITCs protective effect on breast cancer identified in some research, despite the fact that not in other folks, was primarily confined to those possessing the low activity genotype of GSTs. There is certainly suggestive proof that ITC induced apoptosis is mediated by oxidative stresslipid peroxidation items.
We also examined regardless of whether the compounds that most co
We also examined no matter if the compounds that most drastically synergized with cisplatin would sensitize cells to IR. Geldanamycin and SB218078 synergized with IR in both 2008 and 2008 FANCF cells, G?6976 in 2008, and compound 5373662 in 2008 FANCF cells. Other compounds showed additive effects with IR. Discussion Right here we identified 26 chemicals that inhibit the formation of IR and cisplatin induced FANCD2 foci. Quite a few demon strated a stronger inhibition of FANCD2 foci formation than FANCD2 mono ubiquitination, suggesting that at lower concentrations they interfere with FANCD2 recruit ment at web site of DNA harm a lot more than with FANCD2 mono ubiquination. On the other hand, the cathepsinB inhibitor CA 074 Me demonstrated a stronger inhibition of FANCD2 mono ubiquitination than foci formation, sug gesting the intriguing possibility that recruitment of FANCD2 at internet sites of DNA damages may perhaps be supported with decreased levels of mono ubiquitinated FANCD2.
Additionally, selleck inhibitor most chemical compounds also inhibited IR induced RAD51 foci formation and DNA double strand break repair by HR, but frequently not BRCA1 foci formation, indicating that they inhibit multiple discrete kinase inhibitor Paclitaxel methods from the DNA harm response and are certainly not precise inhibitors of your Fanconi anemia pathway. Numerous on the identified chemical substances appeared to cluster around common targets, for instance the proteasome, PKC, CHK1, CDK, HSP90, cathepsin B and lysosome function, or casein kinase II. A few of these targets have already been implicated within the FA pathway and HR. For instance, proteasome function is expected for activation with the FA pathway and HR.
Constant with this, amongst the new FA pathway inhibitors, we identified a novel and uncharacterized proteasome inhibitor. ATR and its downstream kinase, CHK1, which is often straight or indirectly inhibited  by UCN 01, G?6976, SB218078, alsterpaullone, roscovitine and wortmannin, are involved in FA pathway activation. CHK1 inhibi tion also inhibits RAD51 binding to DNA. HSP90 can also be implicated inside the FA pathway and HR, since FANCA, BRCA2, CHK1 and CDKs are clientele of HSP90. CDK inhibition results in perturbation of cell cycle, proliferation and checkpoints, and compromises CHK1, BRCA2 and RAD51 functions, which can cause impaired FA pathway and HR. A doable part for PKC, cathepsin B, lysosome and casein kinase II within the regulation of the FA pathway and HR has not been reported however, and is worth testing in the future. Whether or not these chemical compounds straight target some components with the FA pathway remains to become determined.
by UCN 01, G?6976, SB218078, alsterpaullone, roscovitine and wortmannin, are involved in FA pathway activation. CHK1 inhibi tion also inhibits RAD51 binding to DNA. HSP90 can also be implicated inside the FA pathway and HR, since FANCA, BRCA2, CHK1 and CDKs are clientele of HSP90. CDK inhibition results in perturbation of cell cycle, proliferation and checkpoints, and compromises CHK1, BRCA2 and RAD51 functions, which can cause impaired FA pathway and HR. A doable part for PKC, cathepsin B, lysosome and casein kinase II within the regulation of the FA pathway and HR has not been reported however, and is worth testing in the future. Whether or not these chemical compounds straight target some components with the FA pathway remains to become determined.
Moreover, we showed that this, actually, was the expected result
Moreover, we showed that this, in truth, was the anticipated result in the context of variation in 4E BP1 and eIF4E expression. As an alternative pre dictive marker, we developed assays to estimate one particular with the crucial functional finish points of mTORC1 signalling, eIF4E activity. We identified this estimate to be substantially connected with rapamycin sensitivity in cell culture. It was notable, however, that estimated eIF4E activity was essentially the most considerable predictor of rapamycin sensitivity for eight on the cell lines, when MCF7 cells had been twice as sensitive as predicted by this relationship. MCF7 cells more than express S6K1, on account of amplifica tion of its gene, one explanation for enhanced sen sitivity in MCF7 cells may be that with constitutively higher S6K1 activity, the cells are dependent upon mTOR induced S6K functions for example extra basic translational effects.
In assistance of this, S6K1 more than expression has previously been associated with improved rapamycin sensitivity. over here Importantly, we also examined no matter if estimates of pre therapy eIF4E activity in clinical breast tumours predicted response towards the mTOR inhibitor everolimus. Disappointingly and in contrast to our in vitro work, we found estimated eIF4E activity did not predict response to mTOR inhibition as assessed by transform in tumour cell proliferation. Nonetheless, we did discover that pre therapy eIF4E activity in tumours was substantially linked with substantial alterations inside the expression of eIF4E and its regulators post therapy.
We interpret this to recommend that cancers with higher eIF4E activity might indeed have been sensitive to everolimus, as suggested by our in vitro data, but that the cells remaining right after two weeks of drug remedy reflect choice to acquire drug resistance by changing the order inhibitor pathways regulating eIF4E function. Data show that this proposed resistance will not be necessarily connected with lower estimated eIF4E activity or greater proliferative rates. This hypothesis highlights a difference among quick term sensitivity assays in vitro and longer term drug remedies in patients, inside the latter case it is actually inevi tably extra difficult to assess the early response of tumour cells to remedy and there’s considerable scope for acquired alterations to take location. Ultimately, it truly is exciting to note that our information do not help the use of phospho 4E BP1 as either a predictive or pharmaco dynamic marker for mTOR inhibitors as some have attempted considering the fact that it really is clear that changes in phos pho 4E BP1 relate not only to inhibition of 4E BP1 phosphorylation, but in addition to dramatic modifications in overall 4E BP1 expression. Background Epithelial to mesenchymal transition is usually a biologi cal process in polarized epithelial cells, which occurs in various physiological and pathological conditions.
Whereas the expression of PYGO1 is affected by the well-known TAK
Whereas the expression of PYGO1 is impacted by the well-known TAK1 IKK2 cascade for SLAMF6 and IRF4 also the TAK1 p38 cascade seems to play a function. IgM mediated MYC inhibition is reversed by the PI3K inhibitor Ly294002. This demonstrates an involvement of PI3K signalling to inhibit aberrant MYC expression. In addition, an effect of JNK, IKK2 or PI3K inhibition on basal expression of MYC could be observed. This supports a function of a tonic activation of PI3K, JNK and IKK2 mediated signalling activity in regulating aberrant basal MYC expression. Interestingly, a brand new murine model for lymphomas has been described supporting the view of a synergistic action of c Myc and PI3K signalling. Additionally, a tonic BCR signalling and PI kin ase activity in Burkitts lymphoma has been not too long ago described by Schmitz and co workers.
On the other hand, this link among tonic PI3K signalling and MYC expression has not been described in this publication. Interestingly, in this study therapy of BL lines with BKM120, a PI kinase inhibitor in clinical trials, or rapamycin, an inhibi tor with the mTORC1 complex, was toxic to most BL lines soon after four days. Hence, their rapamycin signature must be taken into account for future selelck kinase inhibitor investigations. Surpris ingly, IKK2 inhibition was connected using a significantly stron ger IgM mediated suppression of MYC expression. As a result, we observed a suppressive part of tonic IKK2 activity onto MYC expression in BL2 cells. This sheds new light onto the regulation of the ab errant expression of MYC.
Good and adverse signals from PI3K, MAPK and NF kB pathways can now be investigated in much more detail by way of example to be able to delin eate variations between BLs and DLBCLs characterized by a higher Myc index or MYC break. A comparable impact of PI3K inhibition as described for MYC is observed also for BCL6, LEF1 and BCL9. How ever, as for MYC, the expression inhibitor NLG919 of BCL6 or BCL9 is currently affected to some extend by Ly294002 in un stimulated BL2 cells. Thus, it’s tough to interpret these data for BCL6 and BCL9 to the end. We speculate that combinations of pathways are involved in both basal and IgM mediated gene expression. In Figure 7A a scheme summarizes the key effects of kinase inhibition observed just after IgM treatment. As already noted above, in some instances the remedy of cells with inhibitors is connected with an enhanced activation or inhibition of respective genes.
For ex ample treatment of cells with Ly294002 led to a stron ger activation of EGR2 or CCR7 by IgM treatment. Comparable effects are observed for IKK2 inhibition for SLAMF3 and ID3, for p38 or JNK inhibition analysing SGK1, ID3 or PYGO1 respectively. In Figure 7B a respective sum mary of most important IgM enhancing effects 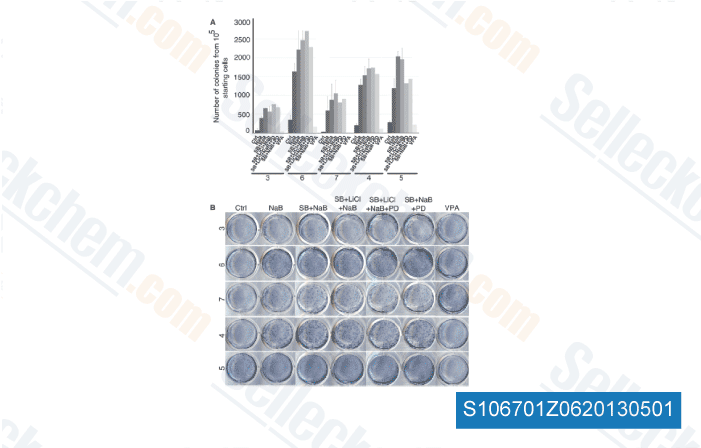 after inhibition of particular kinases is shown, which includes effects of these inhibitors onto the basal expression levels of analysed genes as one example is MYC or BCL9.
after inhibition of particular kinases is shown, which includes effects of these inhibitors onto the basal expression levels of analysed genes as one example is MYC or BCL9.
